Drug development is extremely slow and expensive. The average research and development (R&D) lifecycle lasts 10–15 years, with a significant portion of candidates failing during clinical trials. A major hurdle has been identifying the correct protein targets early in the process.
Proteins are the “functioning molecules” of living organisms – they catalyze reactions, transport molecules and serve as targets for most modern drugs. The ability to rapidly classify proteins, understand their properties, and identify less-researched candidates could dramatically accelerate the discovery process (e.g. Wozniak et al., 2024, nature chemical biology).
This is where the convergence of data engineering, machine learning (ML) and generative AI becomes transformative. In fact, you can build this entire pipeline on a single platform – the Databricks Data Intelligence Platform.
what are we building
Our AI-Powered Drug Discovery Solutions Accelerator Demonstrates end-to-end workflow through four key processes:
- Data Ingestion and Processing: Over 500,000 protein sequences are captured and processed Uniprot.
- AI-powered classification: A transformer model is used to classify these proteins as water soluble or membrane transport.
- Insight Generation: Protein data is rich with LLM-generated research insights.
- Natural Language Exploration: All processed and enriched data is made accessible through AI-enabled dashboards and environments that support natural language queries.
Let’s go through each step:
Step 1: Data Engineering with Lakeflow Declarative Pipelines
Raw biological data rarely comes in a clean, analysis-ready format. Our source data comes like this FASTA Files—a standard format for representing protein sequences that looks like this:
To the untrained eye, it is almost impossible to interpret this sequence data – a dense series of single-letter amino acid codes. Yet, by the end of this pipeline, researchers can query this same data in natural language, asking questions like “Show me less-researched membrane proteins in humans with high classification confidence” And in return gaining actionable insights.
Using Lakeflow declarative pipelines, we build a medal architecture that progressively refines this data:
- Bronze Layer: Raw ingestion of FASTA files, extracting IDs and sequences using BioPython.
- Silver Lining: Parsing and structuring – We extract protein names, organism information, gene names and other metadata using regex transformations.
- Gold/rich layer: Curated, analysis-ready data rich with derived metrics like molecular weight – ready for dashboards, ML models and downstream research. This is the credible layer that analysts and scientists directly inquire about.
Result: Clean, controlled protein data in the Unity Catalog, ready for downstream ML and analytics. Critically, the data lineage that extends beyond this step to other steps (highlighted below) provides incredible value for scientific reproducibility.
Step 2: Protein Classification with the Transformer Model
When it comes to drug discovery not all proteins are created equal. Membrane transport proteins – which are embedded in the cell membrane – are particularly important drug targets because they regulate what enters and exits cells.
we take advantage ProtBERT-BFDA BERT-based protein language model, from RostLab, has been specifically fine-tuned for membrane protein classification. This model treats amino acid sequences like language, learning relevant relationships between residues to predict protein function.
The model outputs a classification (as membrane or soluble) with a confidence score, which we write back into the Unity catalog for downstream filtering and analysis.
Step 3: Enriching Data with GenAI
Classification tells us What Is a protein. But researchers need to know why it matters—What is the recent research? Where are the gaps? Is this an under-explored drug target?
This is where we bring in the LLM. Leveraging both Databricks’ Foundational Model API and external model endpoints, we create registered AI functions that enrich protein records with research context.
Step 4: Natural Language Exploration
We bring everything together in one AI/BI dashboard with Genie Space enabled.
Researchers can now:
- filter protein Based on organism, classification score and protein type
- Explore Delivery Molecular weight and classification confidence
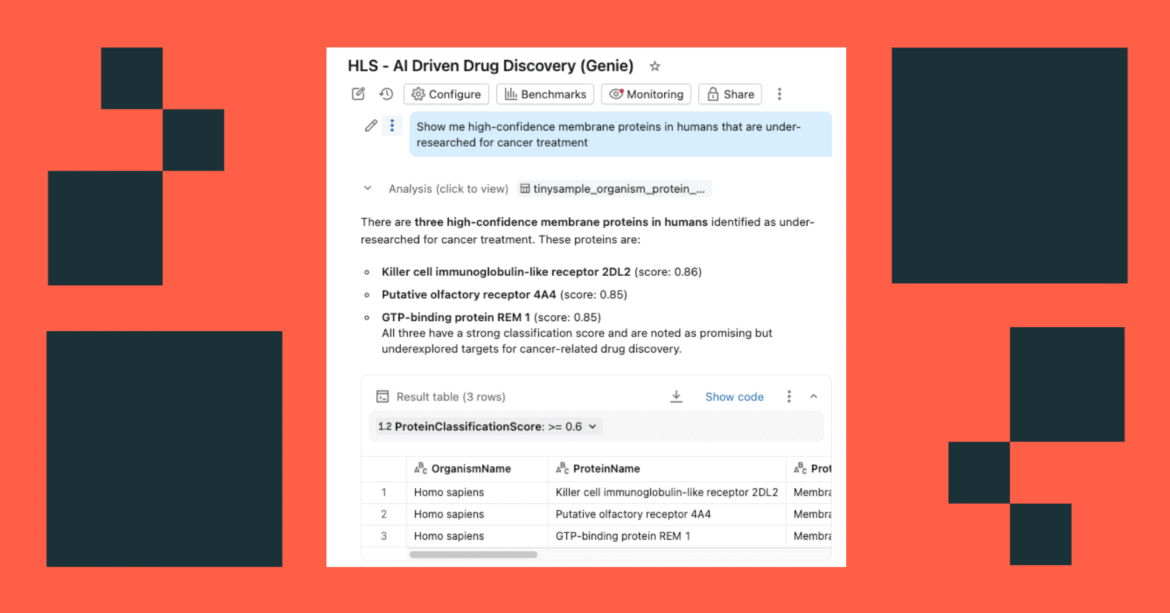
- Ask questions in natural language: “Show me high-confidence membrane proteins in humans that are under-researched for cancer treatment”
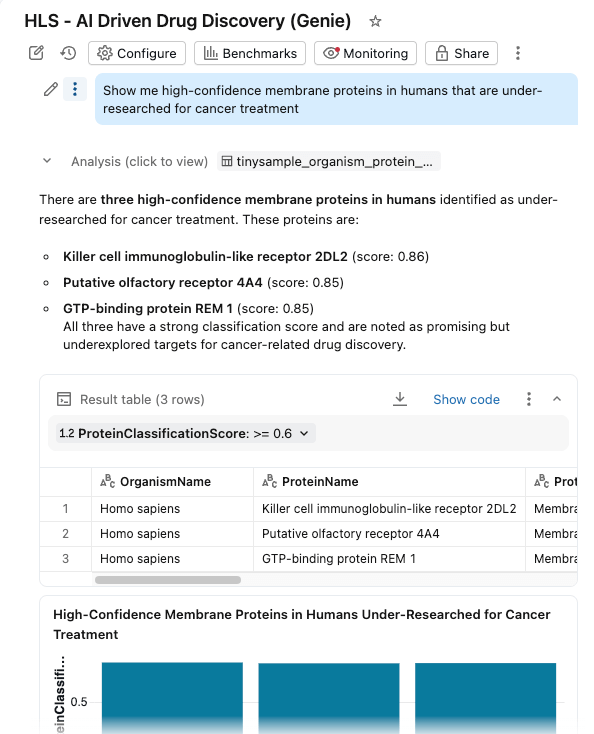
The dashboard questions similar governed tables in the Unity Catalog, with AI functions providing on-demand (or batch processed) enrichment.
The power of a unified platform
What makes this solution attractive is not because of any one component – but rather that everything runs on a single platform:
Critically, there is no data movement between systems. No separate MLOps infrastructure. No disconnected BI tools. The protein sequence that enters the pipeline flows through transformation, classification, enrichment, and becomes queryable in natural language – all within the same governed environment.
The entire solution accelerator is available on GitHub:
github.com/databricks-industry-solutions/ai-driven-drug-discovery
what will happen next
This accelerator showcases the art of the possible. In production, you can extend this to:
- Process the full UniProt database with provisioned throughput endpoints
- Add more (open or custom) classification models for different protein properties
- Construction tile Pipeline on scientific literature for more grassroots LLM responses
- Integrate with downstream molecular simulation workflows
- Connect to protein structure prediction (AlphaFold/ESMfold) to add 3D structural context to classified proteins
- Expand to other genomic formats (FASTQ, VCF, BAM) using Glo for large-scale sequencing and variant analysis
The foundation is there. The platform is integrated. The only limitation is the science you want to accelerate. Get started today.