How close can an open model get to AlphaFold3-level accuracy when it matches the training data, model scale, and estimation budget? ByteDance has introduced Protenix-v1a comprehensive AlphaFold3 (AF3) regeneration For biomolecular structure prediction, released with Code and model parameters under Apache 2.0. model targets AF3-level performance across protein, DNA, RNA, and ligand structures, keeping the entire stack open and extensible for research and production.
Core release also ships with pxmeter v1.0.0An evaluation toolkit and dataset suite Transparent benchmarking on over 6k complexes with Time-division and domain-specific subsets.
What is Protenix-v1?
Proteonics has been described as ‘Proteinics: Protein + X‘, a foundation model for High accuracy biomolecular structure prediction. it predicts All-atom 3D structures for complexes that may include:
- protein
- Nucleic Acids (DNA and RNA)
- small-molecule ligands
The research team defines proteonics as Comprehensive AF3 Reproduction. It reimplements the AF3-style diffusion architecture for all-atom complexes and exposes it in a trainable PyTorch codebase.
The project is released as a full stack:
- training and inference code
- pre-trained model weights
- Data and MSA Pipelines
- a browser-based Protenix Web Server for interactive use
AF3-level performance under matching constraints
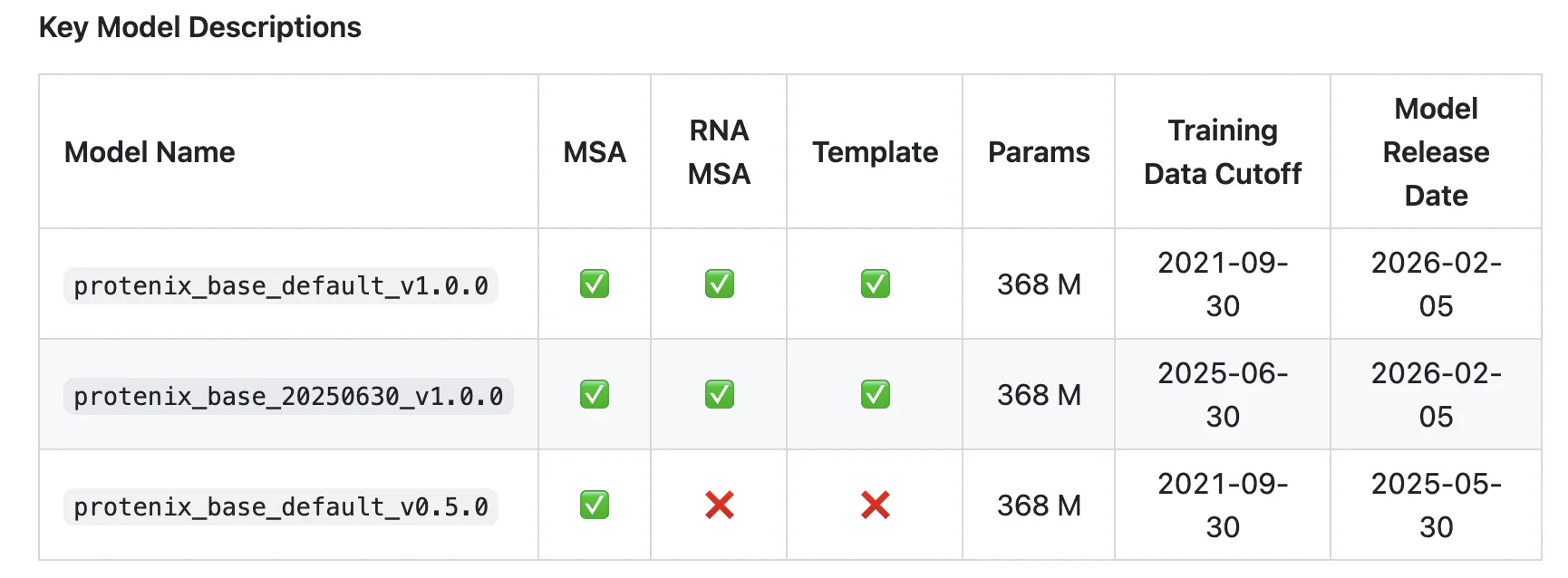
According to the research team proteonics-v1 (protenics_base_default_v1.0.0) Is ‘The first fully open-source model that outperforms AlphaFold3 across a variety of benchmark sets while following the same training data cutoff, model scale, and estimation budget as AlphaFold3.‘
The important obstacles are:
- training data cutoff:2021-09-30, aligned with PDB cutoff of AF3.
- model scale: Protenix-v1 only 368m parameters; AF3 scale has been matched but not disclosed.
- estimated budget: Comparisons use the same sample budget and runtime constraints.
on challenging goals such as antigen-antibody complexis increasing Number of selected candidates from several to hundreds of yields Continuous log-linear improvement in accuracy. It gives a clear and documented information estimation-time scaling behavior Instead of a single fixed operating point.
PXMeter v1.0.0: Evaluation for 6k+ complexes
To support these claims, the research team released pxmeter v1.0.0An open-source toolkit for Reproducible structure prediction benchmark.
PXMeter provides:
- A Manually curated benchmark datasetNon-organic artifacts and problematic entries removed
- Time-division and domain-specific subsets (for example, antibody-antigen, protein-RNA, ligand complexes)
- A integrated assessment framework which calculates complex metrics like LDDT and DOQQ across models
associated pxmeter research papers, ‘Revisiting structure prediction benchmarks with pxmeter,‘ evaluates Proteonics, AlphaFold3, Boltz-1 and Chai-1 on similar curated tasks, and shows how different dataset designs affect model ranking and perceived performance.
How does Proteonics fit into the broader stack?
Proteonics is part of a small ecosystem of related projects:
- pxdesign: A binder design suite built on the Protonix Foundation model. it reports 20-73% experimental hit rates And 2-6× higher success Compared to methods such as AlphaProteo and RFdiffusion, and is accessible through the Proteonics server.
- proteonics-dock: A Classical protein-ligand docking framework Which uses empirical scoring functions instead of deep nets designed for rigorous docking tasks.
- Protenix-Mini And follow up work like Protenix-Mini+: Lightweight variants that reduce inference cost by using architectural compression and few-stage diffusion samples while keeping accuracy within a few percent of the full model on standard benchmarks.
Together, these components cover structure prediction, docking and design, and share interfaces and formats, which simplifies integration into downstream pipelines.
key takeaways
- AF3-class, fully open model:Proteonics-v1 is an AF3-style all-atom biomolecular structure predictor with open code and weights under Apache 2.0, targeting proteins, DNA, RNA, and ligands.
- Strict AF3 alignment for fair comparison: Proteonics-v1 matches AlphaFold3 on significant axes: training data cutoff (2021-09-30), model scale classes, and comparable estimation budget, enabling unbiased AF3-level performance claims.
- Transparent benchmarking with PXMeter v1.0.0: PXMeter provides a curated benchmark suite on 6k+ complexes with time-division and domain-specific subsets and integrated metrics (e.g., complex LDDT, DocQ) for reproducible evaluation.
- Verified Estimate-Time Scaling Behavior: Protenix-v1 shows log-linear accuracy gains as the number of sample candidates increases, giving a documented latency-accuracy trade-off rather than a fixed operating point.
check it out repo And try it here. Also, feel free to follow us Twitter And don’t forget to join us 100k+ ml subreddit and subscribe our newsletter. wait! Are you on Telegram? Now you can also connect with us on Telegram.


